Drug Interaction Solutions is dedicated to supporting scientists in their decision-making process when evaluating pharmacokinetic (PK)-based drug interactions and drug safety.
Drug Interaction Solutions’ PK-based Drug Interaction Database (DIDB®) has the largest manually curated collection of qualitative and quantitative human in vitro and clinical (in vivo) information related to various extrinsic and intrinsic factors. These include interacting co-medications, excipients, food products, herbals, tobacco, organ impairment, and genetics that can affect drug exposure in humans.
You can also find out more in our Why us? brochure.
Data curation
The team of scientists at Certara-Drug Interaction Solutions identify the latest peer-reviewed publications as well as recent NDA/BLA reviews and drug labels from the FDA and select the content that is most relevant to support drug interaction evaluations at various stages of the drug development process. For more information on our data curation process see how we work.
We also create drug monographs that summarize the main mechanistic and quantitative findings including the PK profile, DDI summary, and QT summary, as well as information regarding the overall DDI risk level and label recommendations for clinical use.
In addition, we maintain a DDI Marker Studies Knowledgebase which includes known sensitive and moderate sensitive substrates, weak/moderate/strong perpetrators, based on available clinical evaluations with marker compounds.
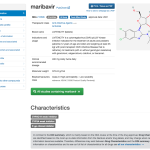
- Monograph for maribavir
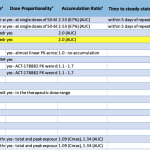
- DDI Marker Studies Knowledgebase – Partial View
Outreach, collaboration, training, and continuous improvement
We share the results of our own research by contributing to workshops and conferences, and publishing articles and reviews on an ongoing basis.
We work closely with colleagues from various universities, regulatory agencies, and pharmaceutical companies on the most pressing issues and challenges in the field.
We assist the end-users of our database with highly specific and detailed database searches and outputs, breaking down often complex mechanisms of drug interactions to enable efficient problem solving. We continuously expand the database content and improve its functionality based on user feedback.