Human in vitro datasets Custom solutions
The clinical datasets contain results from drug interaction, organ impairment, and pharmacogenetics studies.
Common metrics (percent changes in AUC, plasma concentrations, oral and renal clearance values) are used across all studies to allow metadata analysis of quantitative results.
Multiple pre-formulated queries allow users to retrieve an in vivo dataset by drug name, therapeutic class, specific enzyme or transporter, changes in exposure, or toxicity (including QT prolongation).
Detailed study information regarding design, population, dosing of the substrate and the perpetrator, PK, PD, and safety results are extracted from published citations and NDA/BLA reviews and included in all clinical dataset entries.
Comprehensive PK parameters for object (victim) drugs and their metabolites, as well as precipitant (perpetrator) concentrations (when measured) are available.
Results can be viewed, customized, and downloaded in multiple formats, allowing users to compile and organize the large body of information available.
Clinical drug interaction dataset
The clinical drug interaction dataset contains study results from drug-drug, drug-food, and drug-herb, drug-excipient interaction studies and case reports.
Study results are organized according to the overall effect and the underlying mechanism(s) of the interaction:
- Enzyme and/or transport inhibition, induction, or no effect
- Other mechanisms, including absorption-based DDI and food-effect
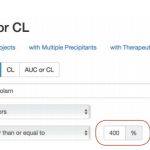
- In vivo interaction query form
- In vivo interaction query results
- In vivo interaction study details
Clinical organ impairment dataset
The clinical organ impairment dataset contains study results from both renal and hepatic impairment studies.
Detailed study information regarding design, population, degree of organ impairment, drug dosing, PK, PD, and safety results are are extracted from published citations and NDA/BLA reviews.
Study results are organized according to the overall effect and the severity of the disease:
- Users can focus on a specific object drug or disease severity (mild, moderate, or severe)
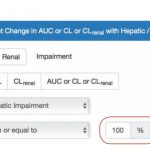
- Explicit changes in exposure can be searched (Example: “Find all drugs with at least 2-fold change in AUC in patients with a specific degree of impairment”)
- Organ impairment query form
- Organ impairment query results
- Organ impairment study details
Clinical pharmacogenetics dataset
The clinical pharmacogenetic (PGx) dataset provides in-depth analysis of the impact of genetic variants of enzymes and transporters on the PK, PD, and safety of drugs in various populations. Available information comes from publications and NDA/BLA reviews describing gene-drug interactions, ethnicity-drug interactions, and case reports.
Detailed study information regarding design, genetic polymorphisms (genotype and phenotype when assigned), and population characteristics are structured and presented according to the latest PGx scientific consensus. This is included in addition to the foundational information of PK/PD, dosing, and safety results included for all clinical study data.
Study results are categorized according to the overall impact of genetic variants on drug exposure, PD, and safety/efficacy compared to a reference group (typically non-carriers of variant).
Comprehensive PK parameters for parent drugs and their metabolites in patients with the variant(s) of interest are available.
- Pharmacogenetics query form
- Pharmacogenetics query results
- Pharmacogenetics study details
Work faster and smarter
What subscribers are using the clinical datasets for:
- PROVIDE CONTEXT for RESULTS OBTAINED with candidate compounds
-
HELP DEVELOP OVERALL REGULATORY STRATEGY and optimize clinical drug interaction trials
- Guide choice of appropriate study design
- Refine inclusion/exclusion criteria
- Help select dose, duration, and timing of drug administration
- Provide PK variability data for power calculations
- Quickly identify known substrates/perpetrators of enzymes/transporters among marketed drugs to understand interaction risk
- SUPPORT STATIC PREDICTIONS and PBPK MODELING
with input parameters - ACCESS REGULATORY DDI STUDIES for recently marketed drugs
- PROVIDE REFERENCE RESOURCE for ASSESSMENT of DRUG SAFETY
- HELP IMPLEMENT PERSONALIZED MEDICINE in the context of pharmacogenetics